Defining and exploiting the indigenous microflora of grapes
Background
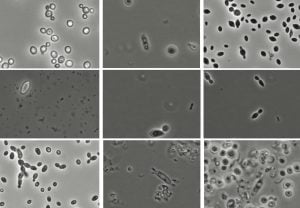
Microorganisms isolated from 2018 winery samples. Photo: K. Sumby
Uninoculated fermentations that use resident grape and wine microflora have seen a dramatic resurgence in winemaking. Previously, fear of spoilage saw resident microbes suppressed by addition of a selected strain and SO2, but now these resident non-Saccharomyces yeasts are being encouraged. The spoilage risk remains, but the reward is an increased flavour complexity arising from extensive competition for and sharing of metabolites and cell-cell interactions. Distinct local populations or ‘microbial terroirs’ have been demonstrated in recent work (e.g Knight et al 2015).
This project will use the unique resource of a single block of many grape varieties on the same soil, encountering the same climatic conditions, to define the impact of grape variety only on microbial terroir. Different varieties, phenology, skin thicknesses, grape and bunch architecture, attractiveness to animal and insect pests, etc are expected to lead to different microbial populations.
Objectives/aims
The project seeks to identify novel yeast and lactic acid bacteria (for use as pure cultures) and an understanding of vine and grape attributes that favour particular species. Knowledge gained about the grape population that inoculates the fermentation will help winemakers better steer the microbes and fermentation to a desired outcome.
Key outputs from this project
Research Articles
KM Sumby, N Caliani, V Jiranek (2021) Yeast diversity in the vineyard: How it is defined, measured and influenced by fungicides. Australian Journal of Grape & Wine Research, 27 (2), 169-193, DOI 10.1111/ajgw.12479
Project leader
Research Associates
Students
Other investigators

Mrs
Kim Chalmers
Chalmers Wines
ON THIS PAGE
Latest News

Macrowine Conference – Isara Vongluanngam
By Isara Vongluanngam Every two years, wine researchers from around the world come together to exchange their latest findings and knowledge at OenoMacrowine conference. This renowned international congress focuses on macromolecules and secondary Metabolites of vine...
Andres Experience at Terclim 2022 | Bordeaux, France
By Andres Zhou Tsang I am sure many of the ARC Training Centre Members have very fond memories of the 13th International Terroir Congress, taking place in Adelaide back in 2020, which connected grape and wine scientists and industry leaders from all over...

Delaying grapes from ripening results in more flavoursome wine
Republished with permission from The University of Adelaide Newsroom Researchers from the University of Adelaide have crunched the data on the best methods to delay grapes ripening on the vine, leading to better quality wine. “Our research focused on three...










